Build, Extend, and Innovate with Platforma
The Platforma SDK is the bridge between raw bioinformatics code and interactive, reproducible workflows — empowering computational teams to build and share production-ready biological applications
Open by design.
Transparent by default.
Inspectable
Zero black-box logic. Dive into the source code of any computational block to perfectly understand how your biological data is being processed and transformed.
Reproducible
Because workflows are entirely code-driven and version-controlled, every analysis run on Platforma is deterministic and fully reproducible across environments.
Community driven
Don't start from scratch. Leverage our expansive library of foundational bioinformatics blocks, or contribute your own.
Built for developers
The core SDK is completely open. Review our architecture, track our release notes, and build with absolute confidence in the underlying framework.
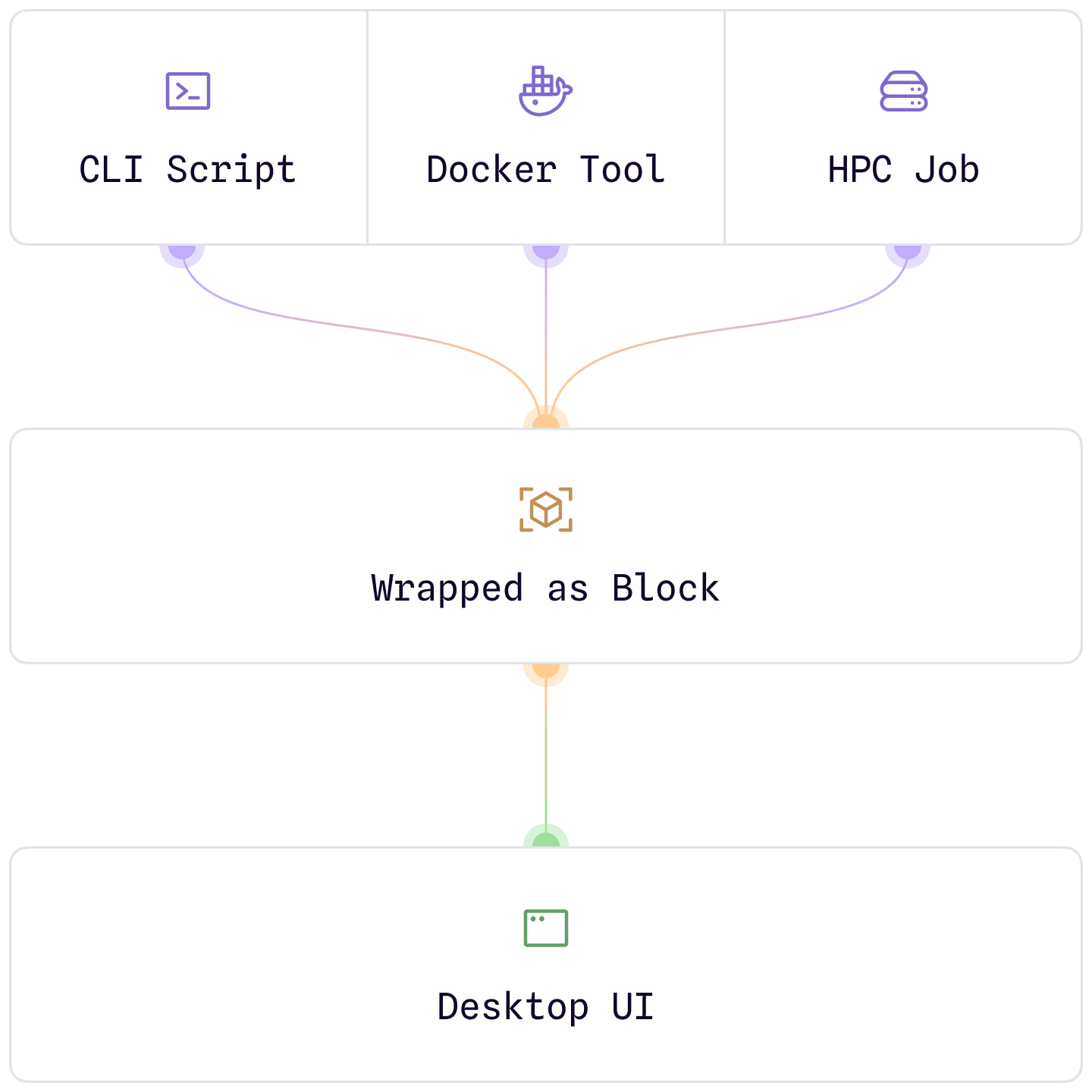
Wrap your pipelines.
Don't rewrite them.
Take your existing command-line pipelines — in-house scripts, Dockerized tools, HPC workflows — and wrap them into interactive, UI-friendly blocks for bench scientists.
No re-engineering. No rewriting in another framework.
Workflow
Define your execution logic using reproducible pipelines. Orchestrate standard command-line tools, capture outputs, and export structured columnar data.
Model
Create the data contract. Map your workflow inputs and outputs into strongly typed, structured arguments and visual components that the UI can consume.
UI
Deliver interactive, intuitive interfaces that run securely inside the Platforma Desktop, giving biologists self-serve access to your pipelines.
Developer experience
Built for BioIT teams
Tooling: Full documentation, local testing environments, and standard block packaging out-of-the-box.
Containerization: Wrap your dependencies securely in containers, ensuring "works on my machine" translates to "works everywhere."
Distribution: Version, publish, and distribute your custom blocks seamlessly via public or private registries.
my block/
workflow/
model/
ui/
test/
package.json
pnpm-workspace.yaml
Security & execution model
Sandboxed UI: The interface runs in an isolated environment with zero unmonitored filesystem or network access.
Controlled runtime: Workflows execute strictly within the protected backend runtime, adhering to your organization's compute and data sovereignty policies.
Immutable: Source-based registry distribution ensures the code you audit is exactly the code you run.