END-TO-END SOLUTION
Multi-omics Analysis
The challenges of
Multi-omics analysis
Disconnected data modalities
Analyzing RNA-seq, proteomics, and immune repertoires in isolation creates a critical analysis gap. Cross-modal insights often require manual export, isolated scripts, and tedious reconciliation.
Scale and dimensionality
As datasets grow massively complex, traditional workflows struggle to fully leverage the data. Without structured dimensionality reduction, meaningful biological signals are easily lost in the noise.
Reproducibility gaps
Relying on fragmented tools, black-box software, and disjointed spreadsheets compromises data provenance. This makes sharing reproducible analyses across cohorts or validation experiments nearly impossible.
Interpretation complexity
Moving beyond simple metrics to uncover "smoking gun" markers and functional relationships between distinct cell states requires advanced, transparent statistical frameworks.
A unified platform
for multi-omics analysis
Platforma provides an integrated computational layer for multi-omics workflows — connecting transcriptomics, immune repertoire data, and experimental metadata into a coherent analytical framework. From raw sequencing data to cross-modal interpretation, Platforma ensures scalable, reproducible, and biologically meaningful analysis.
The Multi-omics analysis suite
Repertoire-transcriptome coupling
Link distinct immune clonotypes directly to their underlying gene expression. Identify relationships between transcriptional programs and clonal expansion. Uncover functional immune responses.
Differential analysis
Quantify transcriptional changes between cell populations. The system integrates the DESeq2 algorithm and interactive Volcano plots to filter noise and isolate robust therapeutic candidates.
Cell type annotation
Leverage automated classification against reference databases or manual tools to refine clusters. Instantly identify uniquely upregulated marker genes to validate cluster identity.
Cohort comparison
Compare patient groups within a robust statistical framework. Distinguish true compositional shifts from simple total cell increases using specialized Bayesian statistical methods.
Functional Analysis
Automatically map differentially expressed gene candidates to established biological pathways, including Gene Ontology and Reactome.

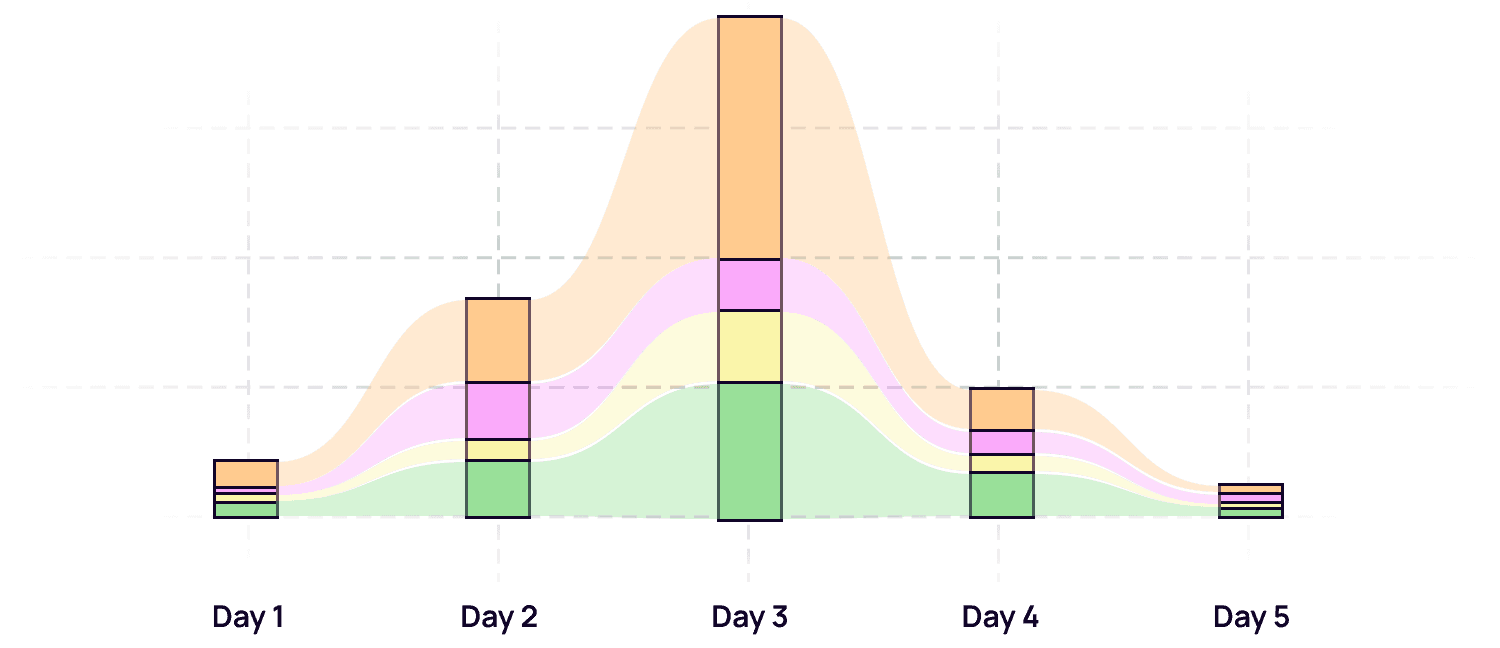
Longitudinal dynamic tracking
Track specific clonotypes across multiple time points or tissues. Visualize the kinetics of immune engagement without complex manual data re-formatting.
Reproducible workflows
Deploy governed, shareable pipelines built on open-source logic. Ensure complete data provenance by tracing every insight back to the raw read.
End-to-end workflow
Application Notes
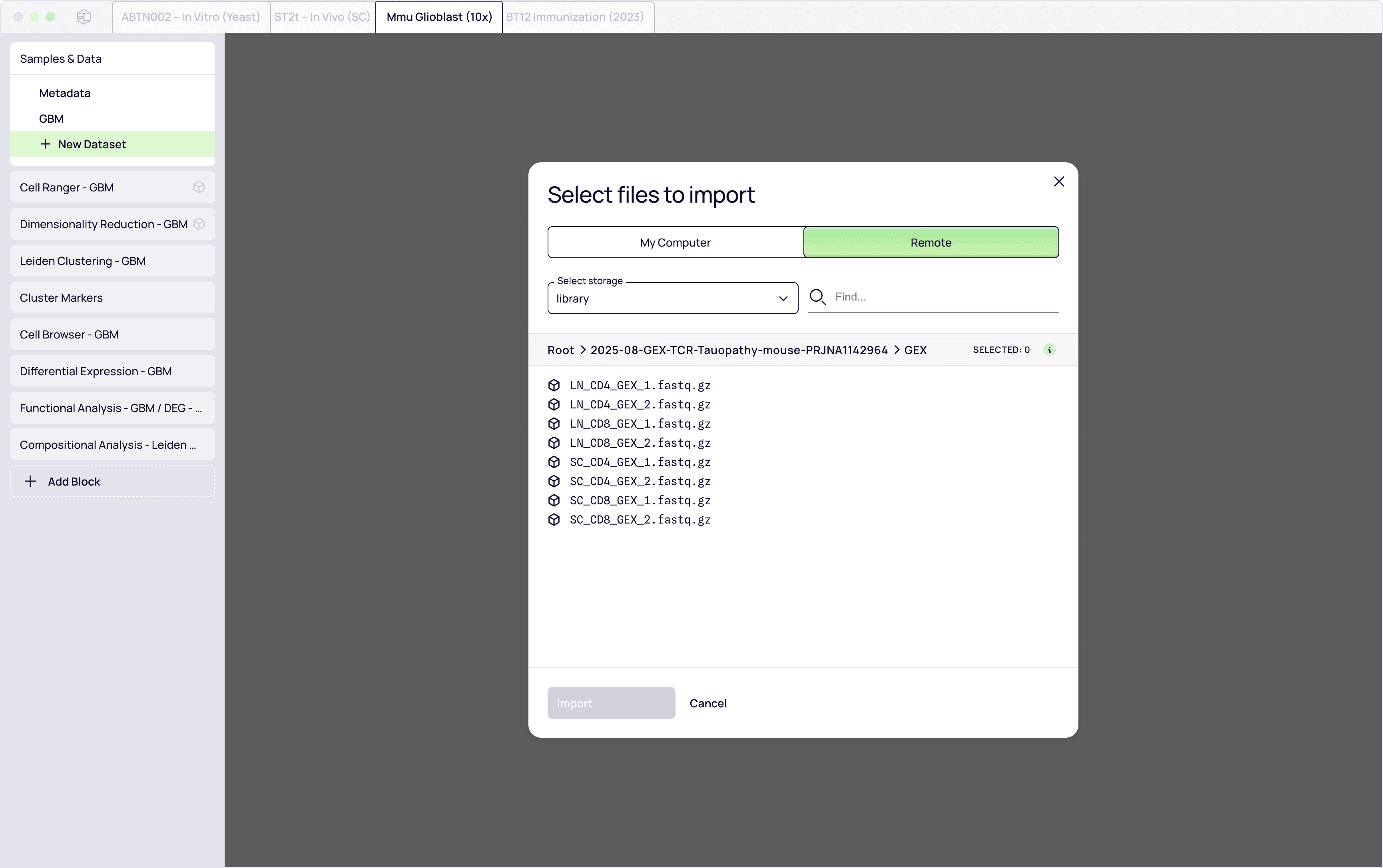
Input data
RNA-seq / scRNA-seq datasets
Immune repertoire sequencing (BCR/TCR)
Clinical and experimental metadata
Preprocessing & QC
Data normalization and filtering
Batch effect correction
Sample validation and harmonization
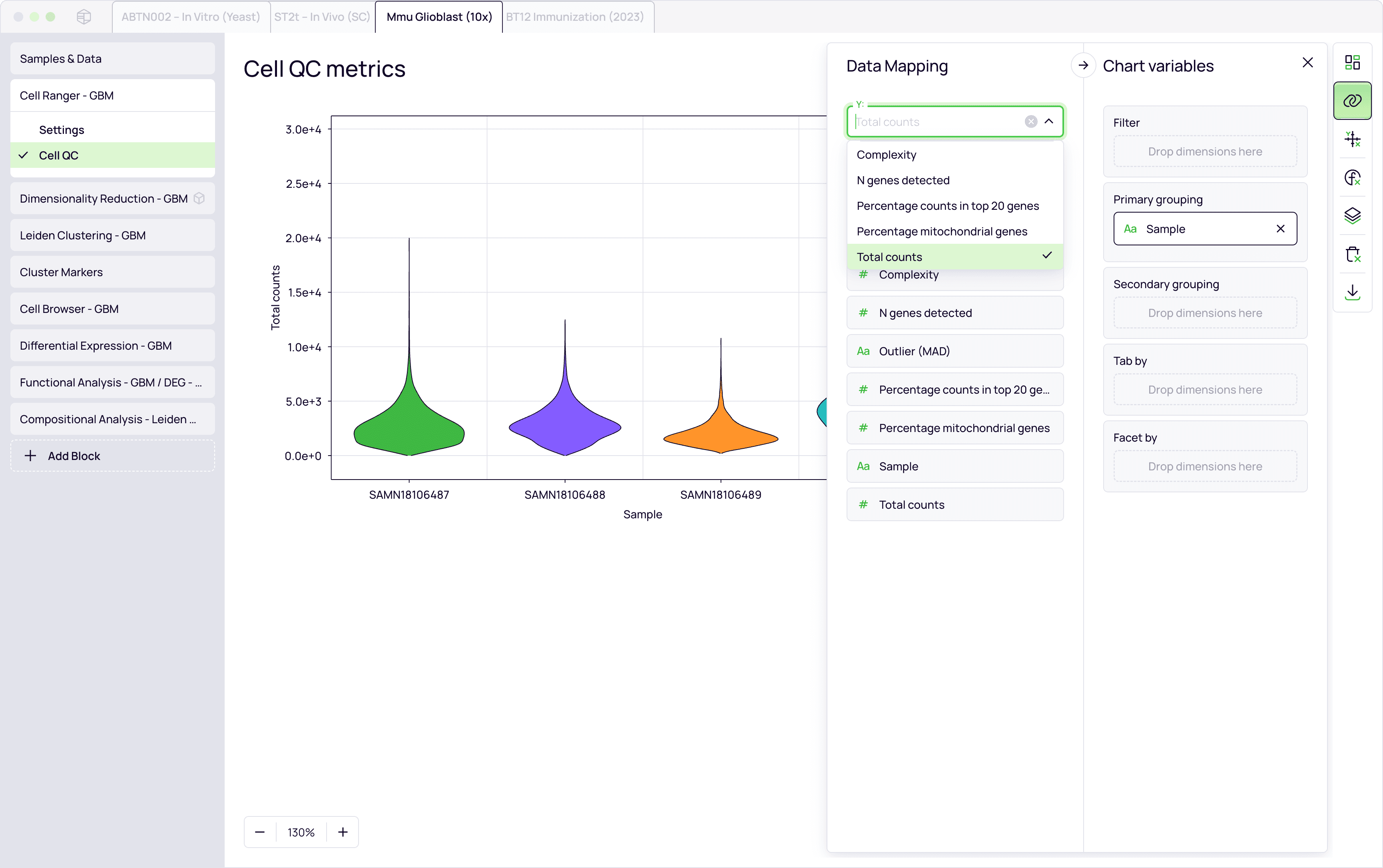
Feature extraction per modality
Gene expression signature profiling
Clonotype identification and diversity metrics
Condition-specific differential analysis
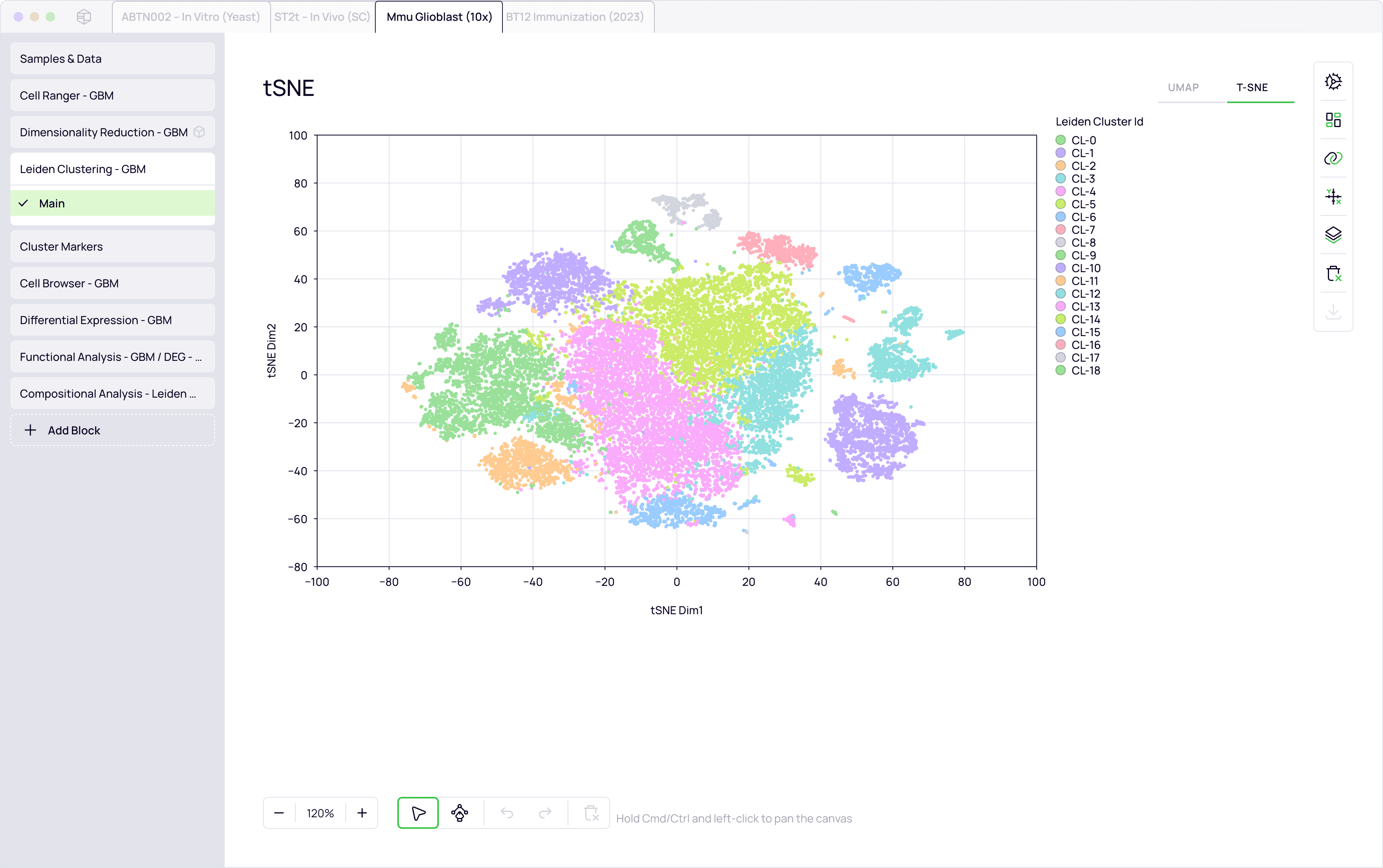
Cross-modal integration
Link clonotypes to transcriptomic states
Dimensionality reduction and clustering
Correlation and enrichment analysis


Longitudinal & Cohort Interpretation
Track immune and transcriptomic dynamics over time
Compare treatment arms or patient groups
Identify coordinated multi-omic response patterns